We are a computational group based in the MRC London Institute of Medical Sciences at Imperial College headed by Boris Lenhard. Our major research interests focus on:
- Structure and function of gene promoters
- Function and genomic distribution of gene regulatory elements
- Function of transcription factors
- Regulation of the production of transcription factors i.e. transcriptional regulatory networks (TRNs)
- Association of different modes of regulation with epigenetic marks and their inheritance
- Transcriptional and translational regulation and dynamics in embryonic development
- The role of core promoters and their diverse architectures in vertebrate development and differentiation
- Emergence and evolutionary dynamics of conserved non-coding elements
Most of our work until now has been purely computational, either analysing publicly available data or in collaboration with experimental laboratories using high-throughput sequencing techniques such as CAGE, ATAC-seq, MNase-seq, RNA-seq, ChIP-seq and Hi-C. Since 2016 we also have experimental activity focusing on studying functional diversity of promoters in development and differentiation.
Visual Abstracts
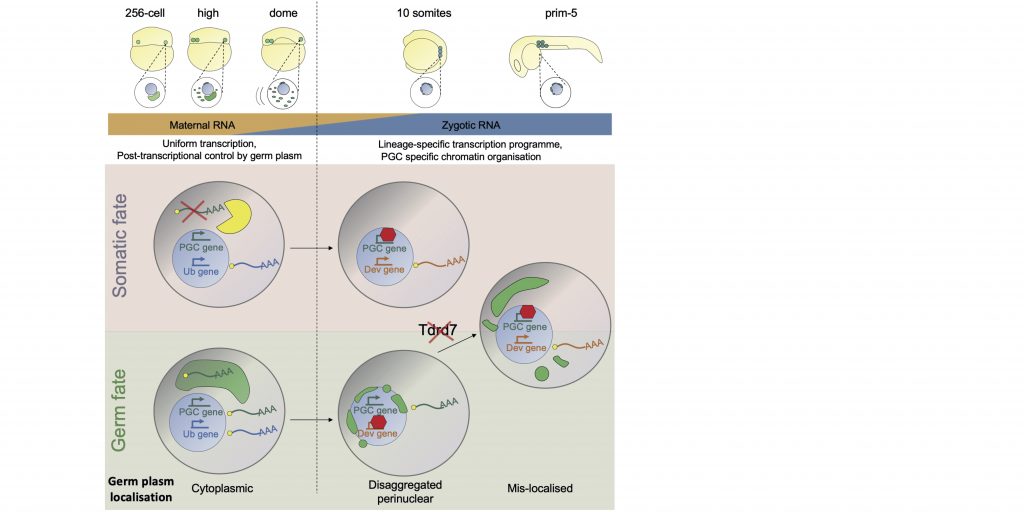
- Germ cell differentiation requires Tdrd7-dependent chromatin and transcriptome reprogramming marked by germ plasm relocalization
https://doi.org/10.1016/j.devcel.2021.02.007
- Global regulatory transitions at core promoters demarcate the mammalian germline cycle
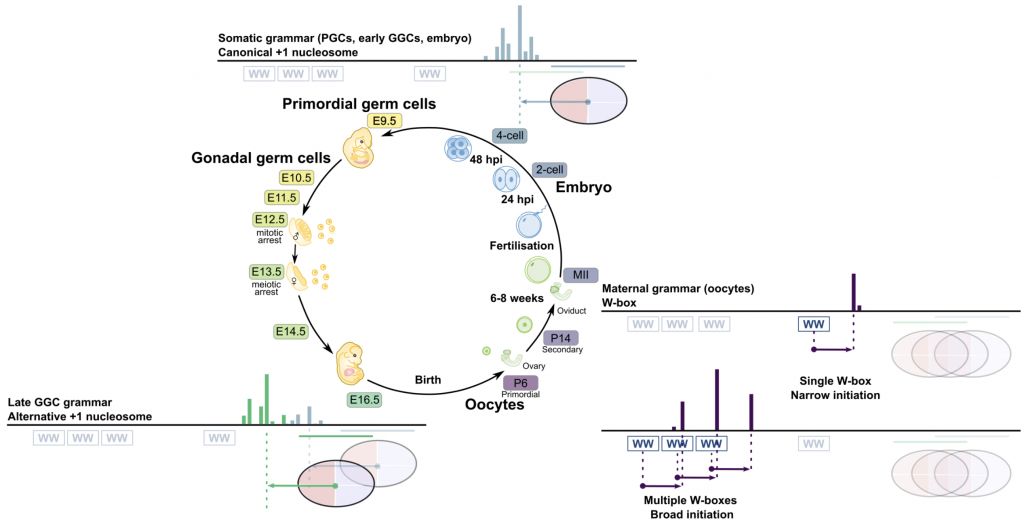
Somatic canonical +1 nucleosome-dependent grammar is used in primordial germ cells, early gonadal germ cells and embryo upon zygotic genome activation. Alternative +1 nucleosome-dependent grammar is used in late gonadal germ cells. Maternal W-box grammar is used in oocytes. Two promoter architecture classes are shown – single W-box leading to a narrow transcription initiation, and multiple W-boxes leading to a broad transcription initiation pattern, distinct from the +1 nucleosome somatic type.
https://www.biorxiv.org/content/10.1101/2020.10.30.361865v1
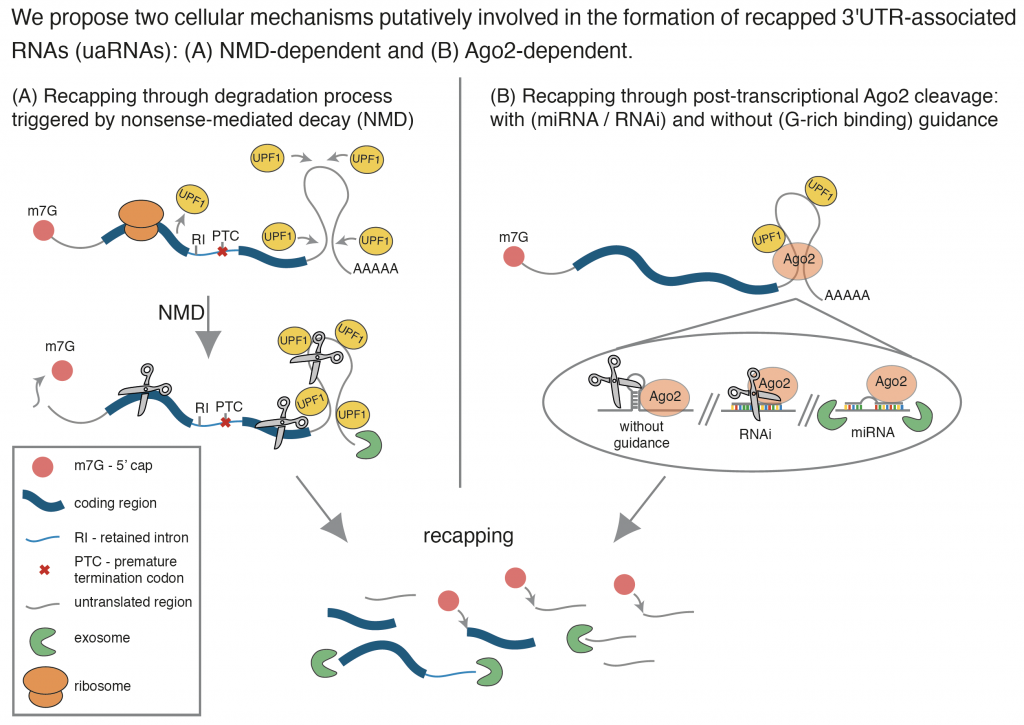
- Transcriptome-wide capping of mRNA fragments